What is Kindler syndrome?
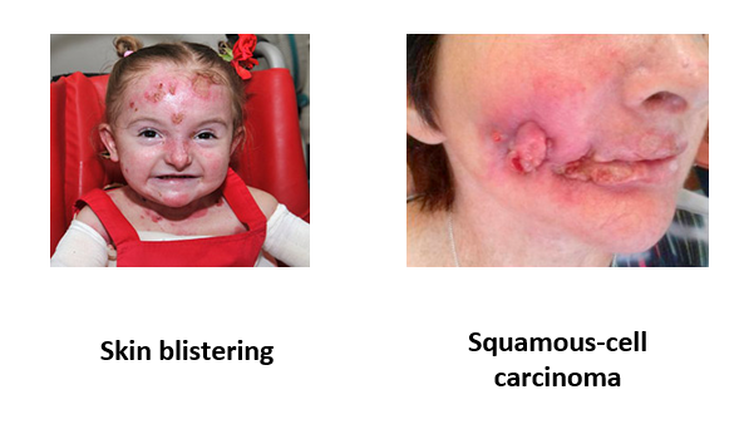
Figure 1: Kindler syndrome phenotypes.
Kindler syndrome is a skin condition that is associated with a variety of different symptoms. The two that will be focused on in this study are skin blistering in young patients and squamous-cell carcinoma development in older patients. An interesting aspect of these two symptoms is that the skin blistering phenotype typically fades around adolescence and the squamous-cell carcinomas are not usually seen in young patients.
|
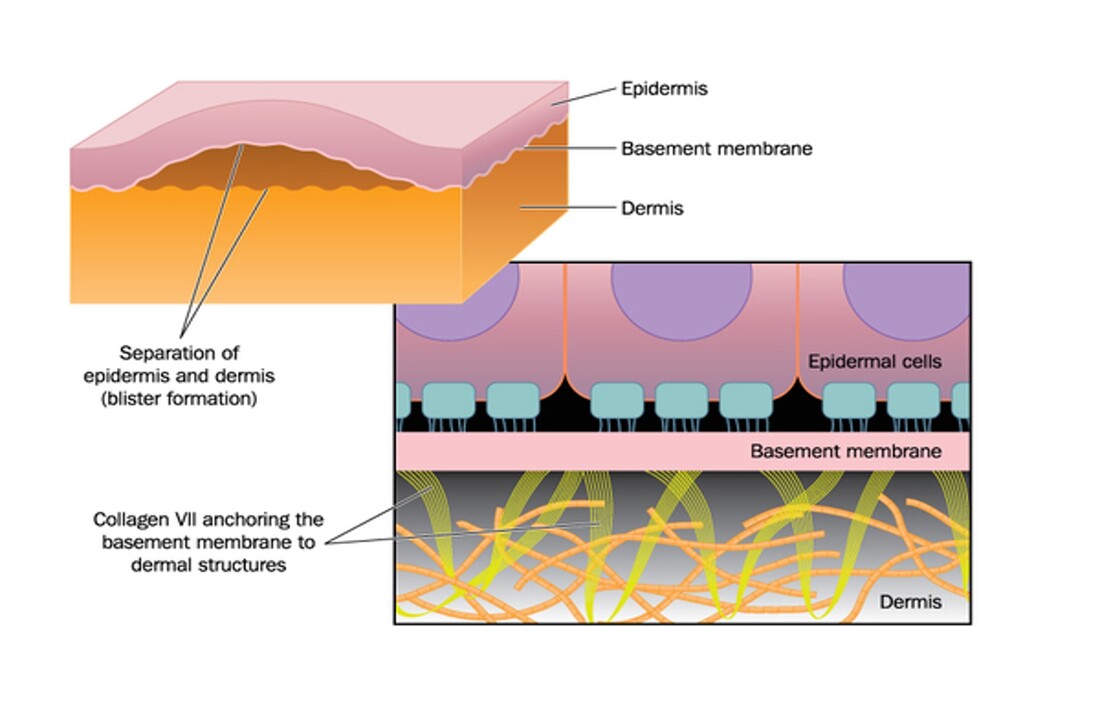
The blistering is caused by a separation of the epidermis and the basal dermis layers of skin. Typically these cells adhere together via extracellular matrix interactions. However, in Kindler syndrome patients, this does not occur. The exact mechanism that causes squamous-cell carcinomas in Kindler syndrome patients is not known beyond the known mechanisms for cancer development. |
Figure 2: Small scale explanation for blistering phenotype
|
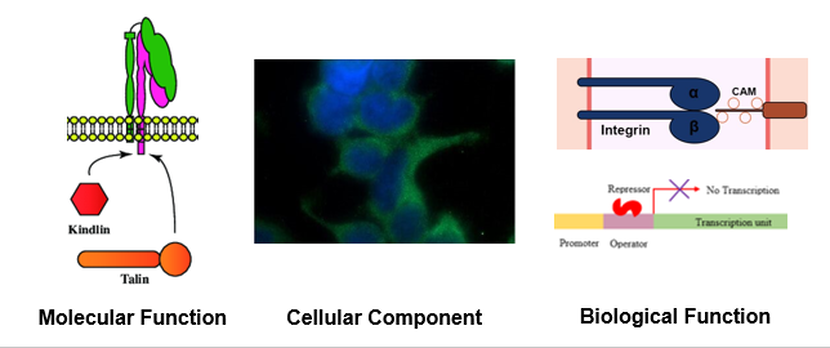
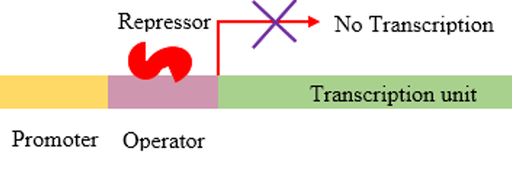
Kindler syndrome is caused by a mutation in the FERMT1 gene, which encodes a protein called kindlin-1. Kindlin-1 functions with talin to activate integrin proteins. It is found in the cytoplasm of cells. Kindlin-1 is involved in the integrin mediated signaling pathway, which leads to the blistering phenotype in Kindler syndrome patients when disrupted, and functions in negative transcription regulation.
Figure 3: Gene Ontology terms for FERMT1
Protein domains and phylogeny
|
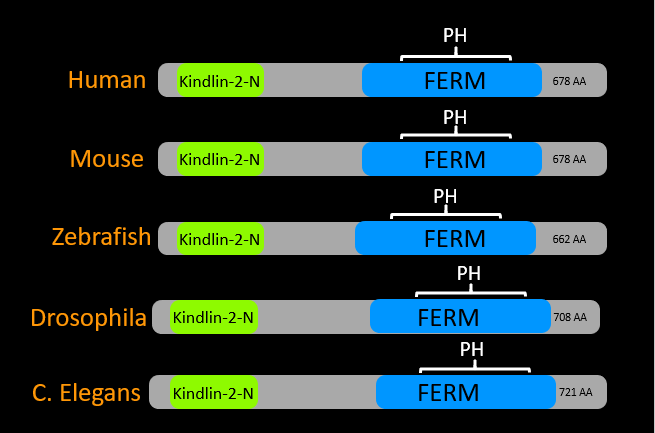
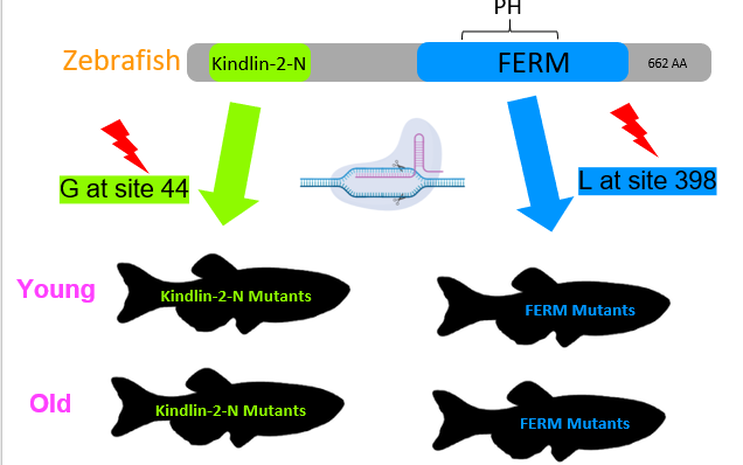
The domains of kindlin-1 are highly conserved among model organisms, including all of its domains. The size of the protein and location of the domains is consistent also. The kindlin-2-N domain is responsible for interactions with talin and integrin proteins. Both the FERM and PH domains serve as structural regions in the protein.
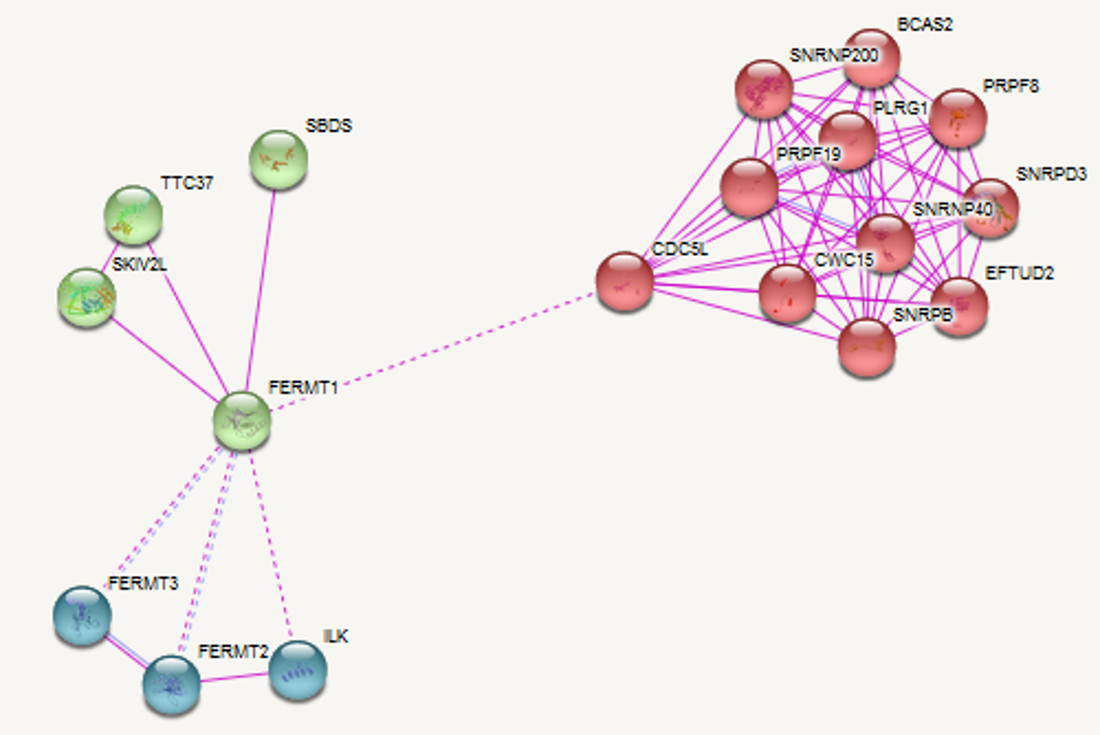
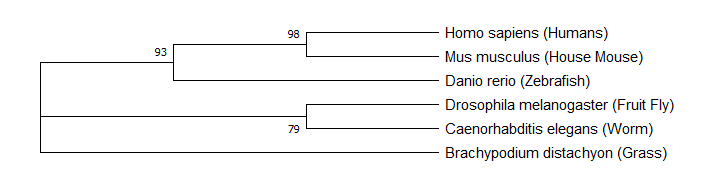
Analysis of the protein interaction network of FERMT1 revealed that it interacts with proteins involved in the integrin mediated signalling pathway (cell adhesion) and transcription related proteins, among others. Specifically, FERMT1's interaction with CDC5L links it to a large network of transcription related proteins. Phylogenetic analysis of FERMT1 revealed that mice and zebrafish FERMT1 are the closest relatives to humans. This is logical because mice and zebrafish have a more comparable skin structure to humans than worms or grass. |
Figure 4: Protein domains of FERMT1 across model organisms
Figure 5: Protein interaction network of FERMT1
|
Figure 6: Phylogeny of FERMT1
Gap in Knowledge
The gap in knowledge for this study is that it is unknown how mutations in FERMT1 lead to cancer development in older Kindler syndrome patients. The hypothesis for this study is that when FERMT1's interaction with transcription proteins is disrupted, overactive transcription leads to uncontrolled proliferation.
Model Organism
|
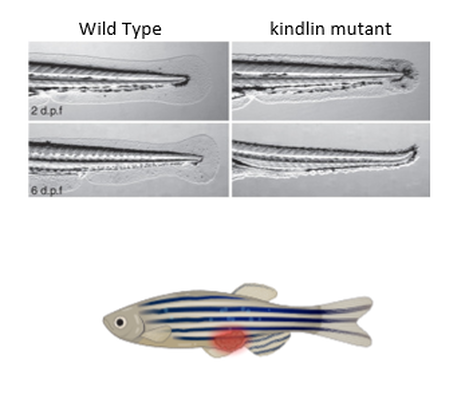
Zebrafish are very well suited to serve as the model organism for this study for a variety of reasons. Firstly, they are significantly less expensive to work with compared to mice. Additionally, kindlin-1 mutants show a phenotypic change over time in the same way that humans do. Young zebrafish show a ruptured fin phenotype that is comparable to the skin blistering phenotype seen in humans. Additionally, older fish have easily observable tumor development. |
Figure 7: (Above) WT zebrafish fins on the left and mutant fins on the right. (Below) A zebrafish with a tumor.
|
Aim 1: Identify conserved residues of fermt1 that maintain cell proliferation
The first test that will be conducted in this study will involve analyzing the aligned sequence of FERMT1 across several model organisms. Analysis of the aligned sequences revealed highly conserved residues within the domains of kindlin-1, a glycine at site 44 within the kindlin-2-N domain and a leucine at site 398 in the FERM domain. These residues will be mutated with CRISPR-Cas9 to truncate the protein and establish mutant lines for this study. Highly conserved residues were chosen because these will be the most likely sites to be involved in cell proliferation since that is a highly conserved process. We expect to see young mutants show the ruptured fin phenotype in youth, but that should fade with age. Also, we expect to see tumor formation in the older mutants, with slightly higher incidence in the kindlin-2-N mutants because more of the protein will be lost in the truncation, increasing the odds of a loss of function of cell proliferation maintenance.
Figure 8: The protein domain of FERMT1 in zebrafish showing where the mutated cites are on the protein and the mutant lines that will be established.
Aim 2: Identify deferentially expressed genes in adult zebrafish mutants vs young
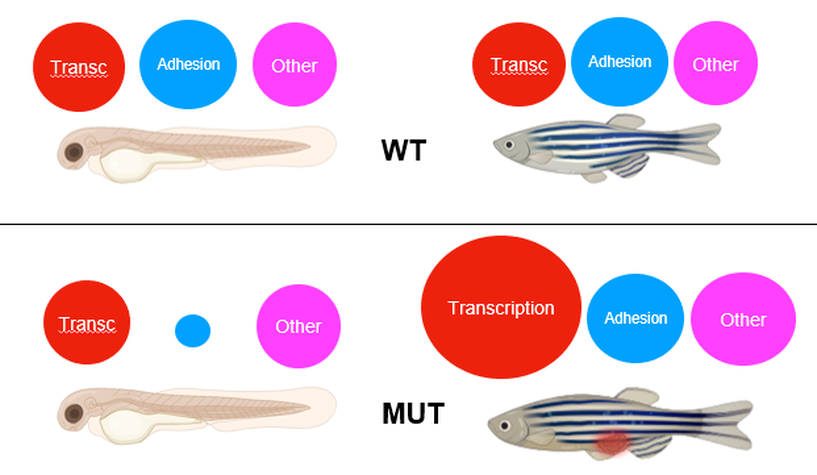
Single-cell RNA-Seq will be performed on cells obtained from the fins of young and adult wild type fish and both established mutant lines. Cells will be taken from the fins because that is the area of the organism that shows the Kindler syndrome phenotype in young fish. We expect to see no change in expression levels between the young and old wild type fish. Both mutant fish are expected to show the same relative expression pattern. It is expected that young mutant fish will show decreased expression of adhesion related genes because FERMT1 will be mutated, and it will explain the ruptured fin phenotype. Adult mutant fish should show increased transcription regulation and adhesion related genes. Transcription related genes expression will be higher than WT and adhesion related genes will return to normal levels.
Figure 9: Red circles represent the relative transcription of transcription related genes. Blue circles represent the relative transcription of cell adhesion genes. Pink circles represent the relative transcription of other genes. (Above) WT young fish and adult fish. (Below) Mutant young fish and adult fish.
Aim 3: identify different protein interaction networks in adult zebrafish vs young
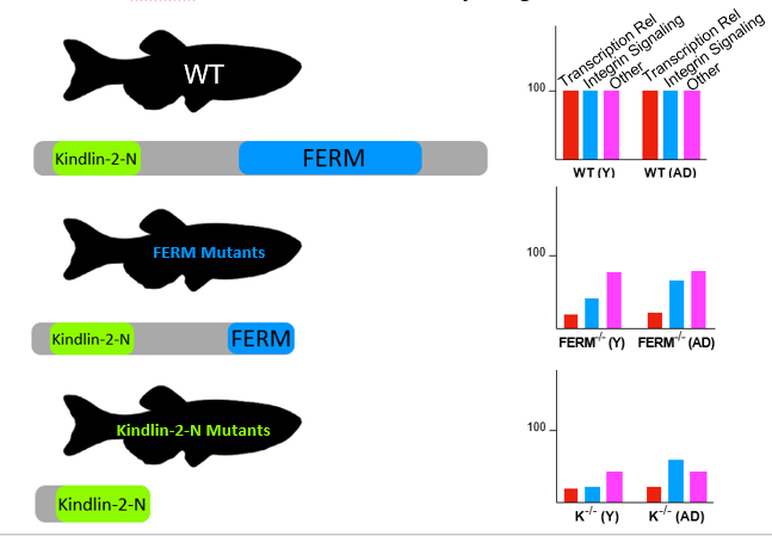
BioID will be used to analyze protein interaction networks in young and old wild type and both mutant zebrafish. BioID will be used because it allows for analysis in vivo. This will allow us to study the same fish over time to ensure any changes seen in the interaction network over time are attributed to aging. Biotin will be attached near the N-terminus before the kindlin-2-N domain to ensure it can bind to the mutant proteins and wild type in the same position. Once again, the wild type fish are not expected to change over time. The FERM mutants are expected to have decreased interaction with transcription related proteins, integrin signaling related proteins, and other proteins in descending intensity, with transcription related proteins showing the largest decrease. Adult fish are expected to recover some integrin signalling protein interactions over time. Young kindlin-2-N mutants are expected to see a drastic decrease in all protein interaction because most of the protein will be lost in the mutation. Again, a small increase in integrin signalling protein interactions is expected in the adult fish.
Figure 10: WT, FERM mutant's, and kindlin-2-N mutant's expected relative interaction with categories of proteins compared to that of WT. WT are 100 as a baseline. Red bars represent the relative protein interaction with transcription related proteins. Blue bars represent the relative protein interaction of cell adhesion proteins. Pink bars represent the relative protein interaction of other proteins.
Future Directions
|
Following this study, it would be interesting to investigate how FERMT2 behaves in similar experiments. Other studies have shown that overexpression of FERMT2 recovers the wild type phenotype in FERMT1 mutants. Investigating if FERMT2 has a role in negative transcription regulation could provide a possible method for treating Kindler syndrome or further insight to the mechanism behind how FERMT1 mutations lead to cancer development. |
Figure 11: FERMT2 recovers function of FERMT1. FERMT1 works as a negative transcription regulator.
|
Final Talk: moyerfinalpresentation592020.pdf
|
REFERENCES
1. Kim, Dae In, and Kyle J. Roux. “Filling the Void: Proximity-Based Labeling of Proteins in Living Cells.” (September 22nd, 2016). Trends in Cell Biology.
2. “Gene Ontology Overview.” Gene Ontology Resource. Retrieved 5/9/2020 from geneontology.org/docs/ontology-documentation/.
3. “Talin (Protein).” (4 May 2020). Wikipedia. Retrieved 5/9/2020 from en.wikipedia.org/wiki/Talin_(protein).
4. “Fermitin Family Homolog 1.” AmiGO 2. Retrieved 5/9/2020 from amigo.geneontology.org/amigo/gene_product/UniProtKB:Q9BQL6.
5. Lowe, Rohan, et al. “Transcriptomics Technologies.” PLOS Computational Biology. May 18th, 2017.
6. Rao, Mohan S., et al. “Comparison of RNA-Seq and Microarray Gene Expression Platforms for the Toxicogenomic Evaluation of Liver From Short-Term Rat Toxicity Studies.” Frontiers in Genetics. January 22nd, 2019.
7. Yandell, Kate. “An Array of Options.” (May 31st, 2015). The Scientist Magazine, Retrieved 5/9/2020 from www.the-scientist.com/lab-tools/an-array-of-options-35381.
8. Kindler Syndrome. (2020, January 21st). Retrieved February 4th, 2020, from https://ghr.nlm.nih.gov/condition/kindler-syndrome.
9. Zheng, Steven, and Ting-Fung Chan. “Chemical Genomics: A Systematic Approach in Biological Research and Drug Discovery.” Current Issues in Molecular Biology. 4 Apr 2002.
10. Dang, Michelle, et al. "Cancer and Zebrafish: Mechanisms, Techniques, and Models: Chemical Genetics." Advances in Experimental Medicine and Biology. 11 May 2016.1. Kindler Syndrome. (2020, January 21st). Retrieved February 4th, 2020, from https://ghr.nlm.nih.gov/condition/kindler-syndrome.
11. Kindler Syndrome. (2016, November 30th). Retrieved February 4th, 2020, from https://rarediseases.info.nih.gov/diseases/4391/kindler-syndrome.
12. Linfante, A. (2018, August 24th). Kindler Syndrome. Retrieved February 6th, 2020, from https://emedicine.medscape.com/article/1118967-overview13. Souldi, H. (2018, February). Kindler syndrome complicated by invasive squamous cell carcinoma of the palate. Retrieved February 6th, 2020, from https://www.sciencedirect.com/science/article/pii/S1879729617300972?via%3Dihub
14. Weber, C. & Ponting, C. "Genes and Homology." (May 4th, 2004). Accessed 5/9/2020 from https://www.cell.com/current-biology/comments/S0960-9822(04)00287-8
15. “Using Research Organisms to Study Health and Disease.” National Institute of General Medical Sciences. Accessed 5/9/2020 from www.nigms.nih.gov/education/fact-sheets/Pages/using-research-organisms.aspx.
16. Mullan, A. & Marsh, A. “Common Model Organisms Used in Molecular Biology.” (February 2019). Oxford Instruments. Accessed 5/9/2020 from andor.oxinst.com/learning/view/article/common-model-organisms-used-in-molecular-biology.
17. Postel, Ruben, et al. “Kindlin-1 Mutant Zebrafish as an In Vivo Model System to Study Adhesion Mechanisms in the Epidermis.” (September 2013). Journal of Investigative Dermatology.
18. Baldauf, Sandra L. “Phylogeny for the Faint of Heart: a Tutorial.” (June 2003). Trends in Genetics.
19. Delsuc, Frédéric, et al. “Phylogenomics and the Reconstruction of the Tree of Life.” (May 1st, 2005). Nature Reviews Genetics.
20. “What Are Protein Domains?” (20 July 2016). EMBL-EBI Train Online. Accessed 5/9/2020 from www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification/what-are-protein-domains.
21. "FERMT1 Search Results" Accessed 5/9/2020 from http://pfam.xfam.org/family/PF00373.18, http://pfam.xfam.org/family/PF18124.1, http://pfam.xfam.org/family/PF00169.29.
22. “Pleckstrin Homology Domain.” (8 May 2020) Wikipedia. Accessed 5/9/2020 from en.wikipedia.org/wiki/Pleckstrin_homology_domain.
23. Kim, Dae In, and Kyle J. Roux. “Filling the Void: Proximity-Based Labeling of Proteins in Living Cells.” Trends in Cell Biology.
22 Sept. 2016. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5077660/
24. "Human FERMT1 Interaction Network." Accessed 5/9/2020 from https://string-db.org/cgi/network.pl?taskId=3vX57XqMI4Oh
25. "Zebrafish fermt1 Interaction Network." Accessed 5/9/2020 from https://string-db.org/cgi/network.pl?taskId=WXDTN8fBpni5
26. “Overview of Post-Translational Modification: Thermo Fisher Scientific - US.” Thermo Fisher Scientific - US. Retrieved 5/9/2020 from www.thermofisher.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/overview-post-translational-modification.html.
27. Helmenstine, Anne Marie. “What Phosphorylation Is and How It Works.” (August 15th, 2019) ThoughtCo. Retrieved 5/9/2020 from www.thoughtco.com/phosphorylation-definition-4140732.
Header image: pngtree.com/free-backgrounds-photos/gene
Epidermis Picture: https://ghr.nlm.nih.gov/condition/kindler-syndrome
Kindler Syndrome picture: https://www.debra.org.uk/uk-funded-projects/sonnenberg-kindler-syndrome
Kindler Knee: https://www.researchgate.net/figure/Clinical-features-of-Kindler-syndrome-a-b-Poikiloderma-with-
hyperpigmentation-and_fig1_229161951
How to treat: https://www.lybrate.com/topic/how-to-treat-kindler-s-syndrome/4dd0715e83fa2628d34f85008f42581e
Worm: http://haasegen564s17.weebly.com/homology.html
Hands: https://www.vectorstock.com/royalty-free-vector/silhouette-hand-helping-hand-vector-631062
Kindler hands: https://link.springer.com/chapter/10.1007/978-3-662-45698-9_43
Zebrafish: https://www.google.com/url?sa=i&url=https%3A%2F%2Fgtgc2016.sciencesconf.org%2Fconference%2Fgtgc
2016%2FYvesClement_2016.07.01_GTGC.pdf&psig=AOvVaw3OSzzPEah2nDhbn3ibyZrP&ust=1582740105733000&source
=images&cd=vfe&ved=0CAMQjB1qFwoTCIiz08Ck7ecCFQAAAAAdAAAAABBP
Kindlin-1 function: https://www.researchgate.net/figure/Molecular-mechanisms-for-integrin-activation-Integrins-exist-in
-two-activation-states-on_fig4_323191947
Zebrafish embryo: http://sitn.hms.harvard.edu/art/2014/zebrafish-embryo-development/
Zebrafish adult: http://www.sleepreviewmag.com/2019/10/zebrafish-study-sheds-light-sleep-regulated-brain/
Squamous Cell Carcinoma: https://www.sciencedirect.com/science/article/pii/S1507136716300517
Biotin: https://en.wikipedia.org/wiki/Biotin
2. “Gene Ontology Overview.” Gene Ontology Resource. Retrieved 5/9/2020 from geneontology.org/docs/ontology-documentation/.
3. “Talin (Protein).” (4 May 2020). Wikipedia. Retrieved 5/9/2020 from en.wikipedia.org/wiki/Talin_(protein).
4. “Fermitin Family Homolog 1.” AmiGO 2. Retrieved 5/9/2020 from amigo.geneontology.org/amigo/gene_product/UniProtKB:Q9BQL6.
5. Lowe, Rohan, et al. “Transcriptomics Technologies.” PLOS Computational Biology. May 18th, 2017.
6. Rao, Mohan S., et al. “Comparison of RNA-Seq and Microarray Gene Expression Platforms for the Toxicogenomic Evaluation of Liver From Short-Term Rat Toxicity Studies.” Frontiers in Genetics. January 22nd, 2019.
7. Yandell, Kate. “An Array of Options.” (May 31st, 2015). The Scientist Magazine, Retrieved 5/9/2020 from www.the-scientist.com/lab-tools/an-array-of-options-35381.
8. Kindler Syndrome. (2020, January 21st). Retrieved February 4th, 2020, from https://ghr.nlm.nih.gov/condition/kindler-syndrome.
9. Zheng, Steven, and Ting-Fung Chan. “Chemical Genomics: A Systematic Approach in Biological Research and Drug Discovery.” Current Issues in Molecular Biology. 4 Apr 2002.
10. Dang, Michelle, et al. "Cancer and Zebrafish: Mechanisms, Techniques, and Models: Chemical Genetics." Advances in Experimental Medicine and Biology. 11 May 2016.1. Kindler Syndrome. (2020, January 21st). Retrieved February 4th, 2020, from https://ghr.nlm.nih.gov/condition/kindler-syndrome.
11. Kindler Syndrome. (2016, November 30th). Retrieved February 4th, 2020, from https://rarediseases.info.nih.gov/diseases/4391/kindler-syndrome.
12. Linfante, A. (2018, August 24th). Kindler Syndrome. Retrieved February 6th, 2020, from https://emedicine.medscape.com/article/1118967-overview13. Souldi, H. (2018, February). Kindler syndrome complicated by invasive squamous cell carcinoma of the palate. Retrieved February 6th, 2020, from https://www.sciencedirect.com/science/article/pii/S1879729617300972?via%3Dihub
14. Weber, C. & Ponting, C. "Genes and Homology." (May 4th, 2004). Accessed 5/9/2020 from https://www.cell.com/current-biology/comments/S0960-9822(04)00287-8
15. “Using Research Organisms to Study Health and Disease.” National Institute of General Medical Sciences. Accessed 5/9/2020 from www.nigms.nih.gov/education/fact-sheets/Pages/using-research-organisms.aspx.
16. Mullan, A. & Marsh, A. “Common Model Organisms Used in Molecular Biology.” (February 2019). Oxford Instruments. Accessed 5/9/2020 from andor.oxinst.com/learning/view/article/common-model-organisms-used-in-molecular-biology.
17. Postel, Ruben, et al. “Kindlin-1 Mutant Zebrafish as an In Vivo Model System to Study Adhesion Mechanisms in the Epidermis.” (September 2013). Journal of Investigative Dermatology.
18. Baldauf, Sandra L. “Phylogeny for the Faint of Heart: a Tutorial.” (June 2003). Trends in Genetics.
19. Delsuc, Frédéric, et al. “Phylogenomics and the Reconstruction of the Tree of Life.” (May 1st, 2005). Nature Reviews Genetics.
20. “What Are Protein Domains?” (20 July 2016). EMBL-EBI Train Online. Accessed 5/9/2020 from www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification/what-are-protein-domains.
21. "FERMT1 Search Results" Accessed 5/9/2020 from http://pfam.xfam.org/family/PF00373.18, http://pfam.xfam.org/family/PF18124.1, http://pfam.xfam.org/family/PF00169.29.
22. “Pleckstrin Homology Domain.” (8 May 2020) Wikipedia. Accessed 5/9/2020 from en.wikipedia.org/wiki/Pleckstrin_homology_domain.
23. Kim, Dae In, and Kyle J. Roux. “Filling the Void: Proximity-Based Labeling of Proteins in Living Cells.” Trends in Cell Biology.
22 Sept. 2016. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5077660/
24. "Human FERMT1 Interaction Network." Accessed 5/9/2020 from https://string-db.org/cgi/network.pl?taskId=3vX57XqMI4Oh
25. "Zebrafish fermt1 Interaction Network." Accessed 5/9/2020 from https://string-db.org/cgi/network.pl?taskId=WXDTN8fBpni5
26. “Overview of Post-Translational Modification: Thermo Fisher Scientific - US.” Thermo Fisher Scientific - US. Retrieved 5/9/2020 from www.thermofisher.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/overview-post-translational-modification.html.
27. Helmenstine, Anne Marie. “What Phosphorylation Is and How It Works.” (August 15th, 2019) ThoughtCo. Retrieved 5/9/2020 from www.thoughtco.com/phosphorylation-definition-4140732.
Header image: pngtree.com/free-backgrounds-photos/gene
Epidermis Picture: https://ghr.nlm.nih.gov/condition/kindler-syndrome
Kindler Syndrome picture: https://www.debra.org.uk/uk-funded-projects/sonnenberg-kindler-syndrome
Kindler Knee: https://www.researchgate.net/figure/Clinical-features-of-Kindler-syndrome-a-b-Poikiloderma-with-
hyperpigmentation-and_fig1_229161951
How to treat: https://www.lybrate.com/topic/how-to-treat-kindler-s-syndrome/4dd0715e83fa2628d34f85008f42581e
Worm: http://haasegen564s17.weebly.com/homology.html
Hands: https://www.vectorstock.com/royalty-free-vector/silhouette-hand-helping-hand-vector-631062
Kindler hands: https://link.springer.com/chapter/10.1007/978-3-662-45698-9_43
Zebrafish: https://www.google.com/url?sa=i&url=https%3A%2F%2Fgtgc2016.sciencesconf.org%2Fconference%2Fgtgc
2016%2FYvesClement_2016.07.01_GTGC.pdf&psig=AOvVaw3OSzzPEah2nDhbn3ibyZrP&ust=1582740105733000&source
=images&cd=vfe&ved=0CAMQjB1qFwoTCIiz08Ck7ecCFQAAAAAdAAAAABBP
Kindlin-1 function: https://www.researchgate.net/figure/Molecular-mechanisms-for-integrin-activation-Integrins-exist-in
-two-activation-states-on_fig4_323191947
Zebrafish embryo: http://sitn.hms.harvard.edu/art/2014/zebrafish-embryo-development/
Zebrafish adult: http://www.sleepreviewmag.com/2019/10/zebrafish-study-sheds-light-sleep-regulated-brain/
Squamous Cell Carcinoma: https://www.sciencedirect.com/science/article/pii/S1507136716300517
Biotin: https://en.wikipedia.org/wiki/Biotin
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison."