What are Protein Domains?
Protein domains are distinct functional or structural regions of a protein. They are the specific parts of the protein that allow it to carry out its functions. Knowing a proteins domains can provide insights to its function and provide valuable information when determining which area of a protein you want to mutate in an experiment. Model organisms can have similar or different domains from a human protein which can affect the results of an experiment.[1]
Pfam and smart databases
There are several different databases that have information on known protein domains. The two used here are Pfam and SMART. Curiously, the SMART database showed different domains for kindlin-1 (the protein produced by FERMT1). The Pfam database showed domains that were supported by primary research articles, so that database was used to construct figure 1.
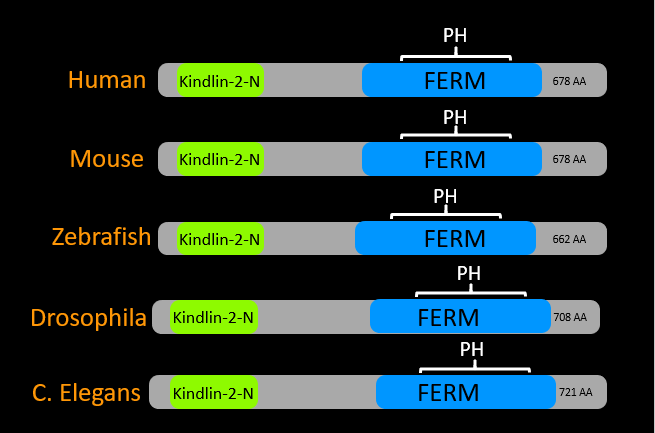
Figure 1: Domains made from data obtained from the Pfam database. Kindlin-2-N domain is in green. FERM domain is in blue and the PH domain is denoted by the white bracket because it is within the FERM domain.
Domain functions
The FERM domain is the central structural domain of the protein. The PH or Pleckstrin homology domain is responsible for intracellular signalling. The kindlin-2-N domain is the N-terminal domain of the protein. This domain is responsible for binding to integrin proteins which are responsible for binding the original cell to other cells. This process is disrupted in Kindler syndrome patients.[2][3]
Discussion
The protein domains of kindlin-1 appear to be highly conserved among model organisms. This would suggest that the protein plays a vital role in a highly conserved pathway. It also shows that a wide variety of model organisms could be used to research issues related to FERMT1 mutations. Since there are no extra domains or domains missing within the protein among model organisms, the protein will likely function similarly across the organisms.
References
Header Image: en.wikipedia.org/wiki/Pleckstrin_homology_domain
1. “What Are Protein Domains?” (20 July 2016). EMBL-EBI Train Online. Accessed 5/9/2020 from www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification/what-are-protein-domains.
2. "FERMT1 Search Results" Accessed 5/9/2020 from http://pfam.xfam.org/family/PF00373.18, http://pfam.xfam.org/family/PF18124.1, http://pfam.xfam.org/family/PF00169.29.
3. “Pleckstrin Homology Domain.” (8 May 2020) Wikipedia. Accessed 5/9/2020 from en.wikipedia.org/wiki/Pleckstrin_homology_domain.
1. “What Are Protein Domains?” (20 July 2016). EMBL-EBI Train Online. Accessed 5/9/2020 from www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification/what-are-protein-domains.
2. "FERMT1 Search Results" Accessed 5/9/2020 from http://pfam.xfam.org/family/PF00373.18, http://pfam.xfam.org/family/PF18124.1, http://pfam.xfam.org/family/PF00169.29.
3. “Pleckstrin Homology Domain.” (8 May 2020) Wikipedia. Accessed 5/9/2020 from en.wikipedia.org/wiki/Pleckstrin_homology_domain.
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison."