What are Protein Interactions?
Many proteins do not function properly unless they are interacting with another protein. Proteins can have several different functions depending on which protein they are interacting with at a given time. Identifying which proteins interact with each other can establish a protein interaction network. Alterations to this network can provide insights to potential treatments or reveal causes for certain phenotypes.[1]
Protein interaction Network of FERMT1
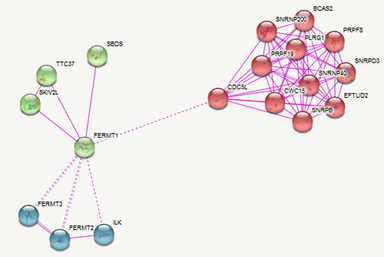
Studies that have conducted experiments to determine protein interactions can have their data submitted to databases. Websites, such as STRING, IntAct, or BioGrid, can utilize this information to build protein interaction networks. The interaction networks established by only experimental data for humans and zebrafish are shown here. In humans, FERMT1 interacts with proteins involved in integrin signalling, translation, and some other processes. In zebrafish, fermt1 interacts with integrin signalling proteins and translation related proteins as well, among others.[2][3]
|
Human
|
Zebrafish
|
|
Figure 1: Protein interaction network for humans. Blue proteins are involved in integrin signalling. Red are involved in transcription. Green are other processes.[2]
|
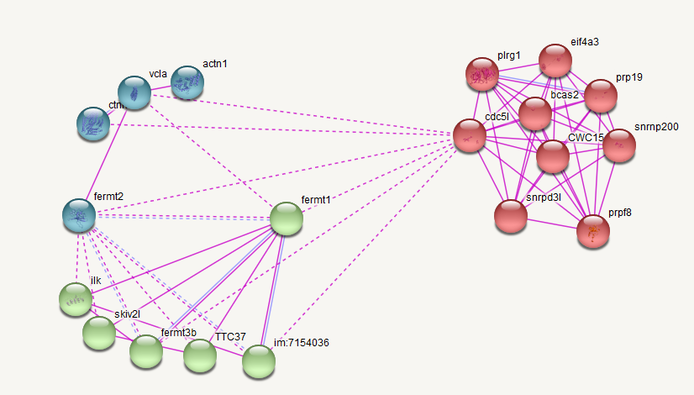
Figure 2: Protein interaction network for zebrafish. Green are integrin signalling related. Red are transcription/RNA processing related proteins. Blue are other processes.[3]
|
Discussion
While the overall interaction network is different for zebrafish, it is expected that zebrafish would have a more complicated interaction network established by experimental data because more assays have been run on zebrafish cells than human. The general processes that the proteins that fermt1 interacts with is fairly consistent between humans and zebrafish. Interestingly, the connection of FERMT1 to translation related proteins, via CDC5L is consistent between the two species and may provide an insight to FERMT1's role in regulating cell proliferation and as a negative transcription regulator.
REFERENCES
Header image: biopharmconsortium.com/publications/advances-in-the-discovery-of-protein-protein-interaction-modulators/
Figure 1: string-db.org/cgi/network.pl?taskId=3vX57XqMI4Oh
Figure 2: string-db.org/cgi/network.pl?taskId=WXDTN8fBpni5
1. Kim, Dae In, and Kyle J. Roux. “Filling the Void: Proximity-Based Labeling of Proteins in Living Cells.” Trends in Cell Biology.
22 Sept. 2016. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5077660/
2. "Human FERMT1 Interaction Network." Accessed 5/9/2020 from https://string-db.org/cgi/network.pl?taskId=3vX57XqMI4Oh
3. "Zebrafish fermt1 Interaction Network." Accessed 5/9/2020 from https://string-db.org/cgi/network.pl?taskId=WXDTN8fBpni5
Figure 1: string-db.org/cgi/network.pl?taskId=3vX57XqMI4Oh
Figure 2: string-db.org/cgi/network.pl?taskId=WXDTN8fBpni5
1. Kim, Dae In, and Kyle J. Roux. “Filling the Void: Proximity-Based Labeling of Proteins in Living Cells.” Trends in Cell Biology.
22 Sept. 2016. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5077660/
2. "Human FERMT1 Interaction Network." Accessed 5/9/2020 from https://string-db.org/cgi/network.pl?taskId=3vX57XqMI4Oh
3. "Zebrafish fermt1 Interaction Network." Accessed 5/9/2020 from https://string-db.org/cgi/network.pl?taskId=WXDTN8fBpni5
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison."